Technical Information
Many reviews of chemical modification of proteins exist.5,6,6a
Some brief examples, extracted from the literature are provided here to illustrate typical
conjugation protocols for surfaces, polypeptides, IgG's, or oligonucleotides.
Part I Conjugation with sulfhydryl reactive probes.
Part II Conjugation with amino reactive probes.
PART I:
Conjugation With Sulfhydryl
Reactive Probes
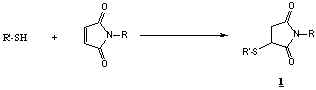
Typical sulfhydryl modification with maleimide:
Use of maleimide derivatives for modification of sulfhydryl groups is described
extensively in the literature5. The maleimide group is specific for
sulfhydryls at ca. pH 6.5-7.5. Reaction with sulfhydryl groups generates a stable
3-thiosuccinimidyl ether linkage (1). The maleimide ring
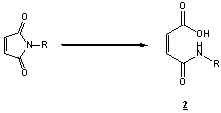
itself can hydrolyze in aqueous buffer to a non-reactive cis-maleamic acid derivative (2)
over long reaction times or extremes in pH (pH>8.0), while competing electrophilically
for free sulfhydryl groups7,8. A study of the rate of
hydrolysis of N-ethylmaleimide via UV absorbtion, in 20 mM phosphate, 20 mM acetate
buffer, pH 7.5, 25 °C revealed a K(decomposition)=1.69 X 10-3
moles/min, with t1/2=410 min or 6.8 h. Optimum derivatization of
sulfhydryl groups with maleimides is usually achieved within 2.0-2.5 h, pH 7.0-8.0.


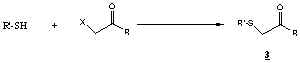
Typical sulfhydryl modification with iodoacetamide:
The use of bromo or iodoacetyl compounds for modification of
sulfhydryl groups on cysteine side chains is described in the literature2,3.
Modification proceedure is similiar to that employed for maleimide, pH 6.5-8.0.
In the absence of free sulfhydryl groups, or those sterically occulded, iodoacetyl
groups can react with the imidazole side chain of histidine9. Reaction
with haloacetyl derivatives should be shielded from light.
Example: Bromoacetyl
modification of BSA4:
To 30 mg of BSA dissolved in 5 mL of 0.1 M NaHCO3, was added 0.2 mL of a
0.7 M solution of Bu3P in 2-propanol. Stir 30 min/25 °C. Add 30 mg
of bromoacetyl derivative, stir 1h. Dialyze for 12 h/4 °C against 0.1 M NH4HCO3
with three changes of same over 2-days. Isolate conjugate by lyophilization.
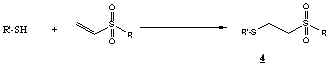
Typical sulfhydryl modification with vinylsulfone:
Vinylsulfone is a slightly softer electrophile than maleimide.
Sulfhydryl addition to vinylsulfones generates a stable b-thiosulfonyl
linkage (4). Sulfhydryl modification with vinylsulfones has
the advantage that the vinylsulfone itself is stable in aqueous solution for days at pH
9.01, thus allowing extended reaction times for modification without
hydrolysis. Sulfhydryl addition to vinylsulfone, unlike maleimides, does not
generate stereoisomers which in some instances may complicate analysis and/or bioactivity
of the conjugate.
Example: Vinylsulfone labeling of RNase1:
RNase (1.35 mg/mL) in 0.2 M, pH 8.0 borate buffer. Vinylsulfone in DMF
(10 mg/mL) stock solution is added to RNase at 22 °C. Reaction is allowed to
proceed until optimum modification is achieved (1-2.5 h). Quench aliquotes with 1.5
M b-mercaptoethanol prior to analysis. Product is
isolated by gel filtration.
Note I: [Reduction of RNase: 14 mg protein (Mr=13,700) dissolved in 300 mL of 25 % b-mercaptoethanol in 4.5 M guanidinium chloride. Heat 100 °C/5 min. The reduced protein (four cystine disulfide bonds reduced to eight cysteine thiol groups) is isolated by gel filtration chromatography using 0.05 M tris, 0.2 M guanidinium chloride pH 4.3.]
Typical sulfhydryl modification with pyridyl
disulfides:
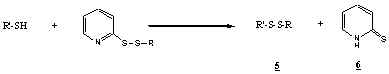
2-Pyridyl disulfide derivatives react with free sulfhydryl groups of
polypeptides or IgG to form disulfide conjugates (5) with release
of 2-pyridylthione (6). Thiolation can be quantitated by
measurement of 2-pyridylthione (lmax=343 nm, e=8.08 X 103 M-1 cm-1).
Conjugates can be cleaved back to their original sulfhydryl groups by addition of excess
free thiols such as DTT, DTE, or BME. Cleavage can also be achieved by use of
non-thiol reducing agents such as Bu3P or TCEP·HCl.
Example: Modification of BSA
with labeled RNase10:
Reduced BSA (80 mg containing 490 nmol of mercaptoalbumin by Ellman's Assay) was
dissolved in 1.95 mL 0.1 M phosphate, 0.1 M NaCl buffer pH 7.5. 2-Pyridyldithio
derivative of RNase (500 nmol) was added in same buffer. Reaction is usually
complete within 2 h, and can be followed by UV measurement of released 2-pyridylthione.
Gel filtration through Sepharose 6B using 0.3 M NaCl provides pure adduct.
For labeling 2-pyridyldithio-small molecule derivatives, dissolve in organic co-solvent if
necessary.
Bibliography
1. Morpurgo, M, Veronese, F.M., Kachensky, D., Harris, J.M
(1996) Bioconjugate Chem. 7, 363-368.
2. Inman, J.K., Highet, P.F., Kolodny, N., Robey, F.A. (1991) Bioconjugate Chem.
2, 458-463.
3. Bernatowicz, M.S., Matseuda, G.R. (1986) Anal. Biochem. 155, 95.
4. Kolodny, N., Robey, F.A., (1990) Anal. Biochem. 187, 136-140.
5. Means, G.E., Feney, R.E. (1990) Bioconjugate Chem. 1, 2-12.
6. Brinkley (1992) Bioconjugate Chem. 3, 2.
6a. Hermanson, G.T. (1996) Bioconjugate Techniques, Academic
Press.
7. Ishi, Y., Lehrer, S.S. (1986) Biophys. J. 50, 75-80.
8. Smyth, D.G., Blumenfeld, O.O., Konigsberg, W. (1964) Biochem. J. 91, 589.
9. Gurnd, F.R.N., (1967) Methods Enzymol. 11, 532-541. See also ref in ref
#5.
10. Carlsson, J., Drevin, H., Axen, R. (1978) Biochem. J. 173, 723-737.
11. Kohno, T., et. al. (1994) unpublished esults with PEG-20K-maleimide.
Additional helpful references: Analysis for extent of sulfhydryl modification.
1. Riddles, P.W., Blakeley, R.L., Zerner, B. (1983) Methods Enzymol.
91, 49-60.
2. Riddles, P.W., Blakeley, R.L., Zerner, B. (1979) Anal. Biochem. 67, 493-502.
3. Ellman, G.L. (1959) Arch. Biochem. Biophys. 82, 70-77.
4. Ellman, G.L. (1958) Arch. Biochem. Biophys. 74, 443.
5.
Technical Assistance:
Additional scientific and technical information on Molecular Biosciences'
products is available on request from our technical service hotline:
Phone: 303.581.7722
FAX: 303.581.0575
Internet: info@molbio.com
Mail: Technical Assistance
Department
Molecular Biosciences, Inc.
4699 Nautilus Court
Boulder, CO 80301
PART II:
Conjugation With Amine Reactive Probes
Typical amine modification with NHS or sulfo-NHS ester:
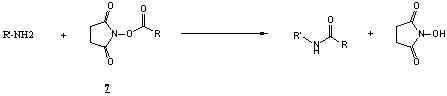
Typical amine modification with isocyanates:
Typical amine modification with isothiocyanates:
Additional helpful references: Analysis for extent of amine
modification.
1. Udenfriend, S., et.al. (1972) Science 178, 871. (Use
of fluorescamine.)
2. (Use of TNBS.)